This post explains how to perform Elastic Net cross-validation (CV) prior to genomic prediction. Elastic Net CV is a well-known machine learning method used to select a subset of SNPs, which helps improve genomic prediction accuracy. Specifically, this procedure iteratively performs CV using Elastic Net across a range of lambda (λ) values. It is important to note that users do not need to manually define these λ values; the software automatically generates and scans them internally.
To perform CVs, the population is repeatedly split into training and validation sets for each λ. While Elastic Net CV is computationally intensive and repetitive, it is a valuable pre-processing step because it can substantially improve the final prediction accuracy.
In order to mitigate the computational burden, QTLmax 6.0 supports GPU acceleration. It is important to note that as of February of 2026, QTLmax 6.0 supports only AMD Radeon GPUs. If a Radeon GPU is not installed in your workstation, you cannot use this function. Supporting NVIDIA CUDA is under consideration, but not ready yet.
Figure 1 shows the “Elastic Net” tab selected in QTLmax 6.0. Unlike other tabs, this tab automatically displays the information about an installed GPU, which is highlighted in blue.
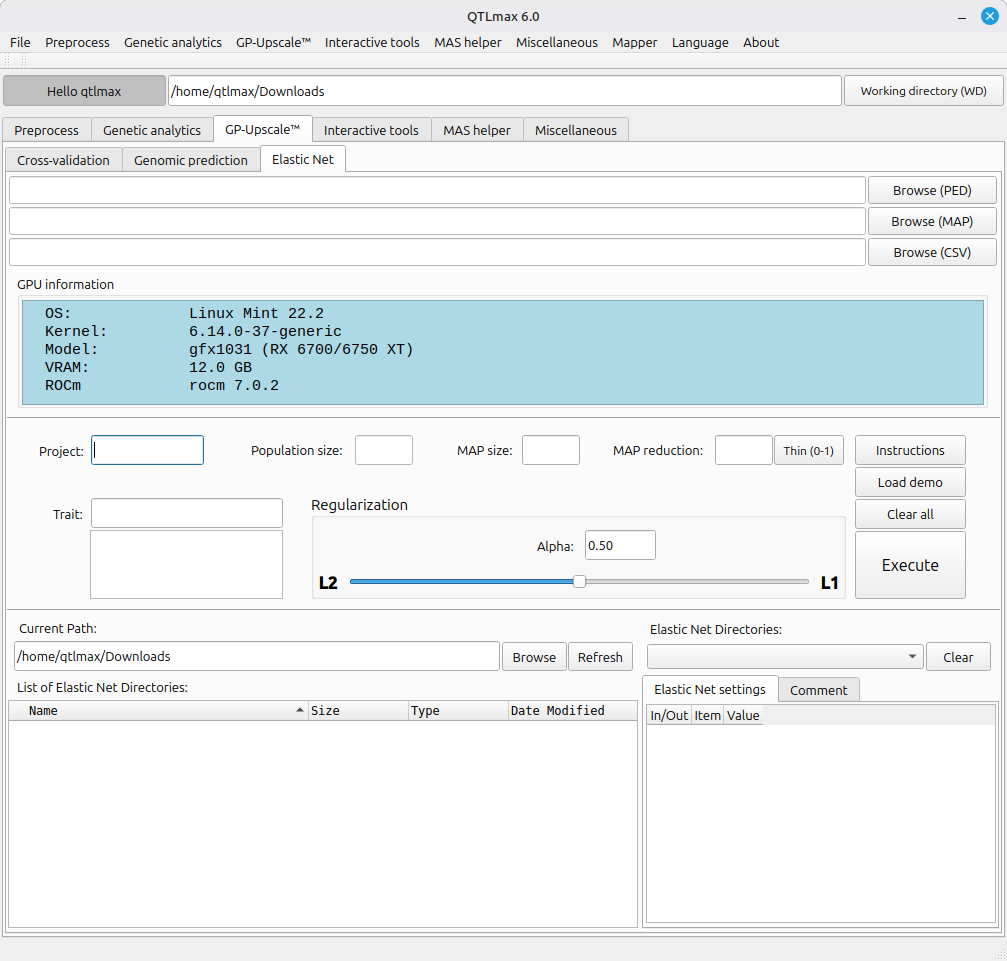
(Figure 1)
Figure 2 shows that once all required input files are chosen, the “Population size” and “MAP size” fields are automatically populated.
Elastic Net requires an Alpha value to be set between L1 (Lasso) and L2 (Ridge Regression). The closer the value is to L1, the fewer SNPs will remain; the closer to L2, the more SNPs will be retained. As an optimal Alpha value must be sought between L1 and L2, the set of SNPs can be refined to achieve the best genomic prediction accuracy. By default, the Alpha value in the Regularization box is set at 0.50. Please note that this is merely a default setting and is not necessarily the most recommended value for every analysis.
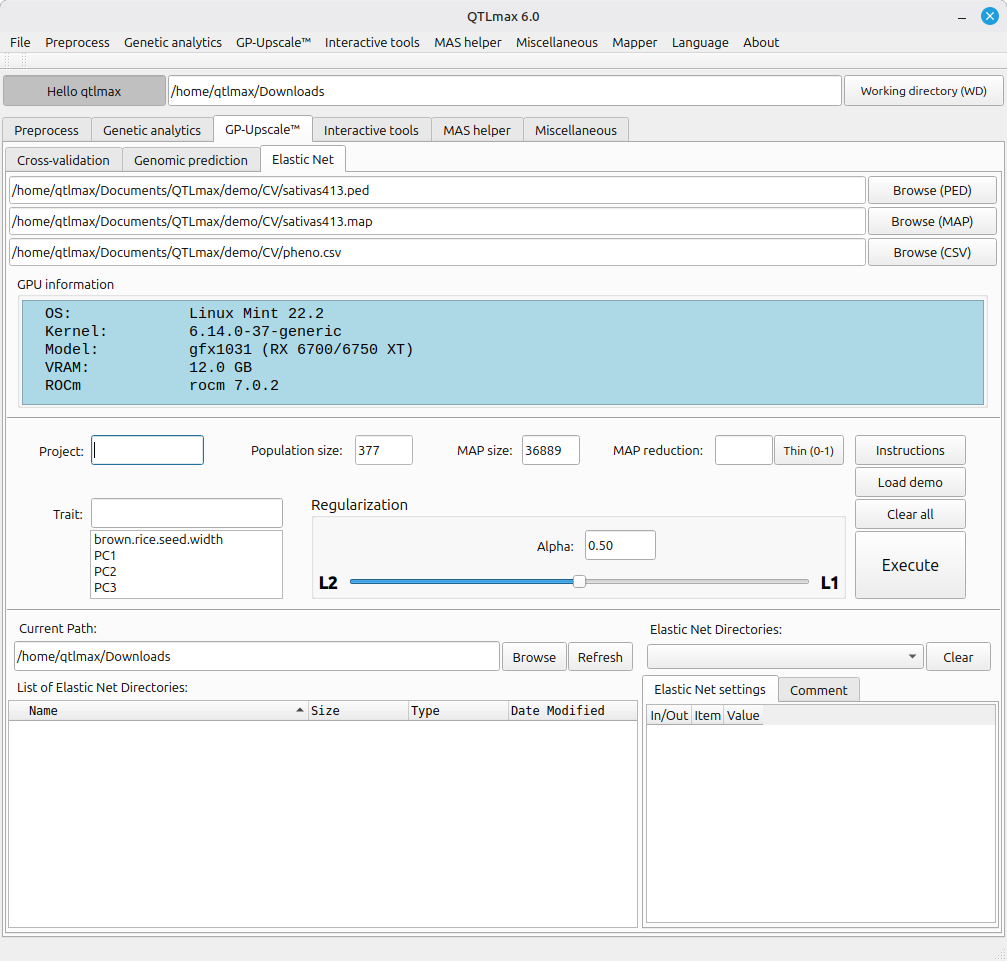
(Figure 2)
Figure 3 illustrates the interface after all required values have been configured and the computation is complete. Upon completion, the project folder (named ‘Enet’ in this example) appears in the bottom-left box. Additionally, the ‘Elastic Net directories’ combo box allows you to select past analysis projects; selecting an item will populate the table with all relevant configuration, input, and output data.
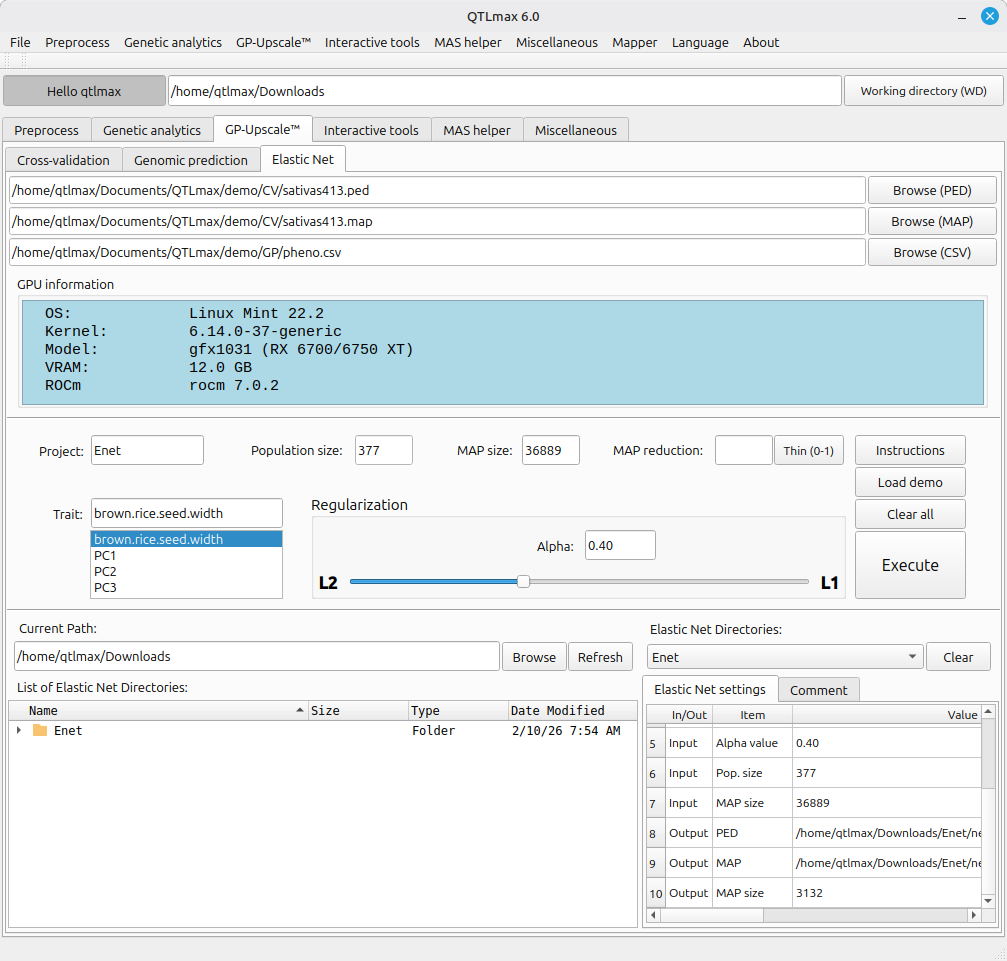
(Figure 3)
As Elastic Net may require multiple iterations with varying values for Alpha, you may want to attach some comment on iteration. To do so, select the ‘Comment’ tab in the bottom-right section, enter your notes in the text box, and click the [Save] button. Once your note is saved, whenever you select the project directory, its note will be retrieved.
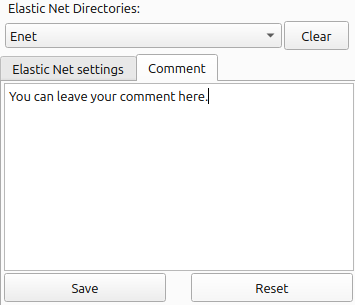
(Figure 4)
It is important to note that results may vary slightly due to the stochastic nature of cross-validation, even with identical inputs. Therefore, it is good practice to perform multiple runs to identify a consensus set of SNPs, thereby ensuring more robust genomic prediction accuracy.