This post explains how to calculate a kinship matrix based on plant/animal SNP dataset. There are many kinds of kinship matrices; although they are intended for measuring genetic relatedness based on its own foundational reasoning, each kind of method has different numeric ranges in resulting outputs.
While the type of kinship matrix addressed in this post is constructed based upon an SNP dataset, it is in the same context with the numerator relationship matrix (NRM); it means its numeric outputs range between 0 and 2.
Traditional NRM only required pedigree records. Once upon a time, before the advent of genomic fingerprinting technologies, pedigree records were the only available data for computing genetic relatedness among entries. These days, we can calculate the high-resolution NRM by taking advantage of SNP datasets, which is the exact topic of this post.
Figure 1 shows the “Genomic Kinship Matrix” tab, where the input dataset has been loaded and the resulting computation is complete; its configuration is literally simple. All requirements are (1) choosing a *.ped file, (2) entering a project name, and (3) selecting the number of CPU threads for this computation.
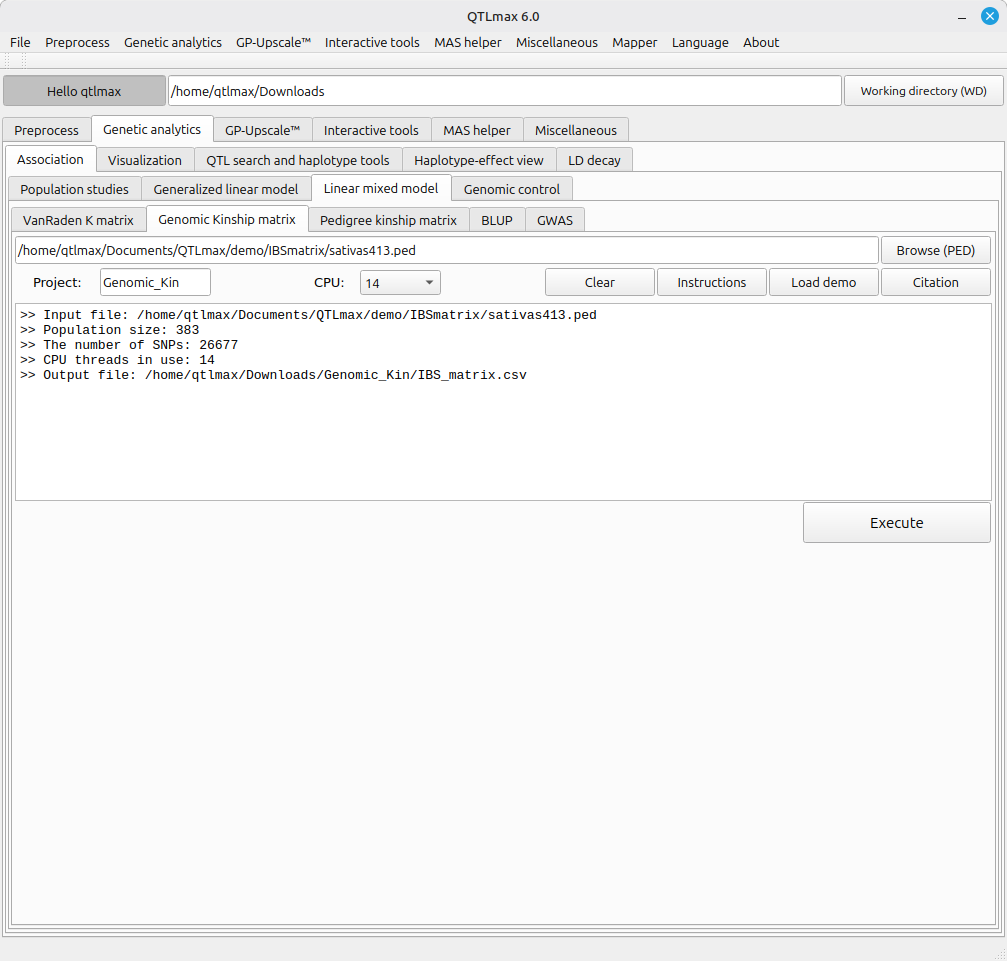
(Figure 1)
Figure 2 shows the resulting genomic NRM based on the given dataset. Suppose that this represents kinship relatedness among entries in a base population, you can simulate a future NRM by expanding this matrix based on scheduled pedigrees. For details, please click on the following link: https://open.qtlmax.com/guide/index.php/2026/02/09/plant-pedigree-kinship-matrix/
Importantly, this simulation can work out because this genomic NRM has the same range as the traditional NRM based on pedigree records. The simulated kinship matrix can be used as an input for ssGBLUP (Single-step Genomic BLUP) in genomic prediction.
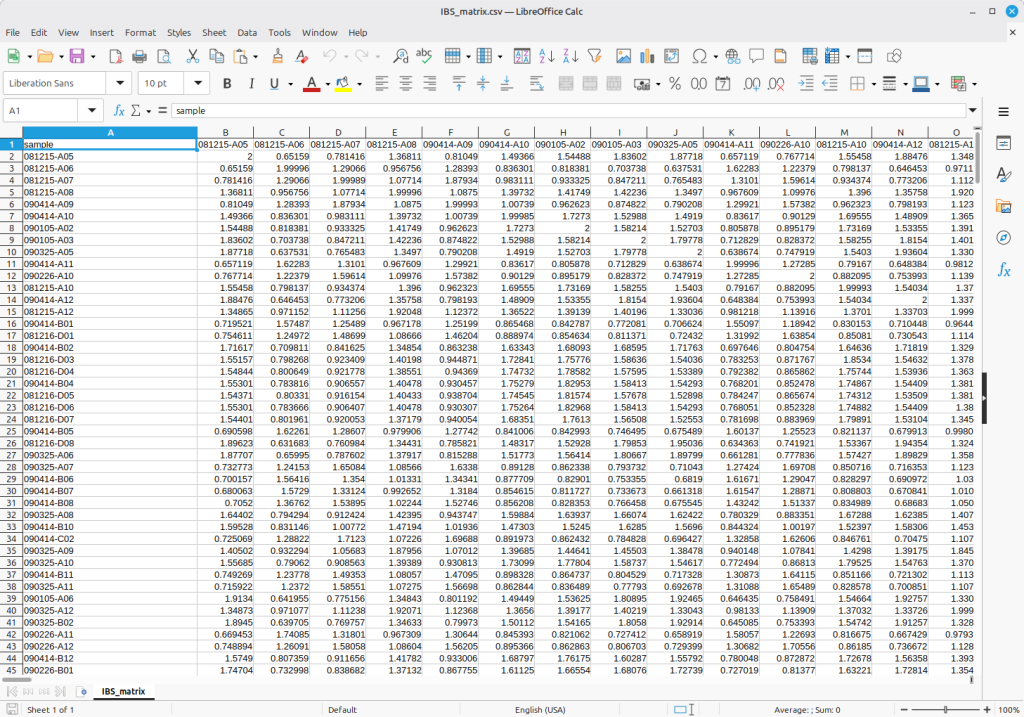
(Figure 2)