This post explains how to calculate a kinship matrix based on plant/animal pedigrees. There are many kinds of kinship matrices; although they are intended for measuring genetic relatedness based on its own foundational reasoning, each kind of method has different numeric ranges in resulting outputs.
The type of kinship matrix addressed in this post is the numerator relationship matrix (NRM) so that the numeric outputs necessarily ranges between 0 and 2. The NRM was dominantly used in linear mixed model frameworks until the advent of genome-based kinship matrices.
Although today’s trend in quantitative genetics mainly relies on genomic kinship matrices, the NRM is still useful. For example, given a genomic kinship matrix derived from a base population, a future kinship matrix can be simulated by applying scheduled mating schemes—represented as pedigrees—to the existing genomic data; the NRM algorithm is essential for this computational process; to this end, of course, the genomic kinship matrix must be constructed on the same scale as the NRM to ensure their numeric ranges are properly aligned. If you want to construct a genomic kinship matrix in the NRM scale, please click on the following link:
Figure 1 shows that a pedigree kinship matrix has been computed in QTLmax 6.0. In this example, “ped.txt” file had been used; as it was loaded, a bunch of pedigree records in the file were displayed in Pedigree editor. Notably, the pedigree notation format systematically follows its own unique syntax. The rules of the syntax were designed based the plant-pedigree format, which can be understood as follows:
Entry name $ ((parent1 / parent2) / parent3)^n
Where Entry name is the progeny’s name; parents 1, 2, and 3 are the progenitors; and n represents the n-th filial generation. It is important to note that each mating event (indicated by a ‘/’) must be enclosed in parentheses to maintain the correct pedigree structure.
The related pedigree notation syntax and its computational algorithm are disclosed via the following publication: https://academic.oup.com/jhered/article-abstract/107/7/686/2622949
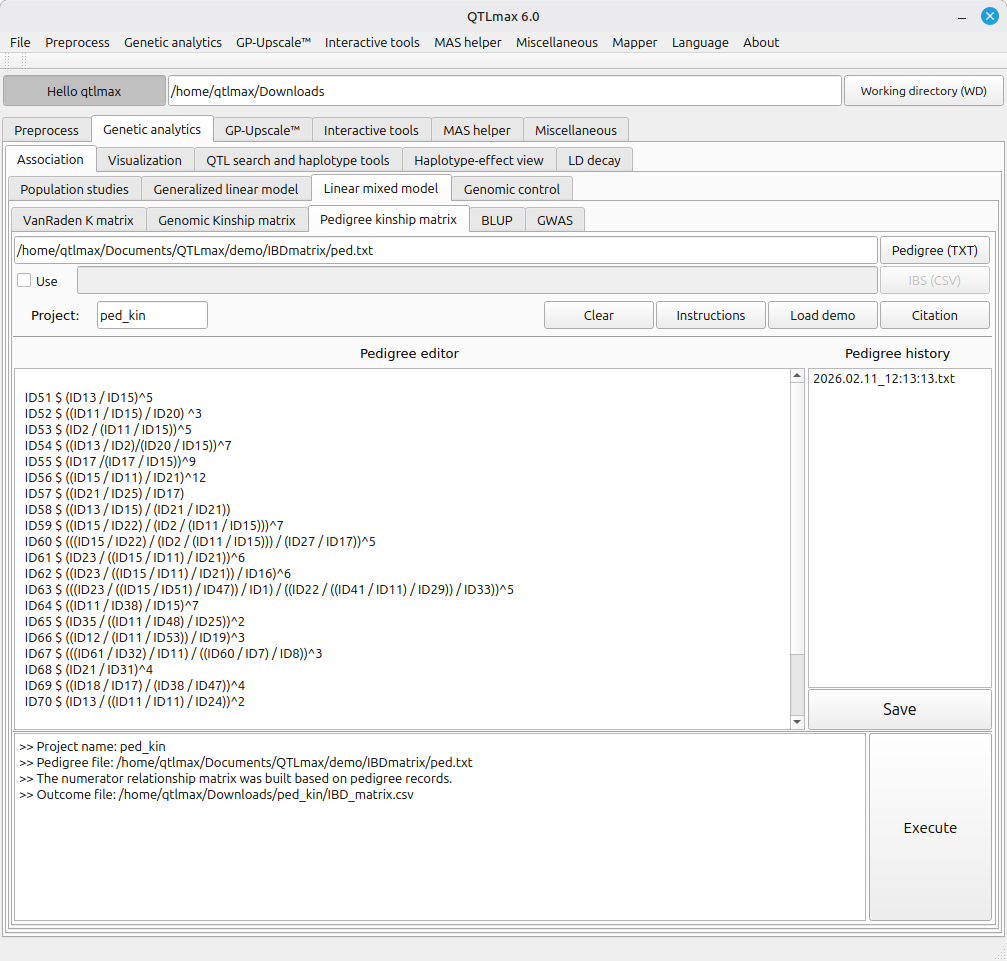
(Figure 1)
Shown in Figure 2 is the resulting pedigree K matrix.
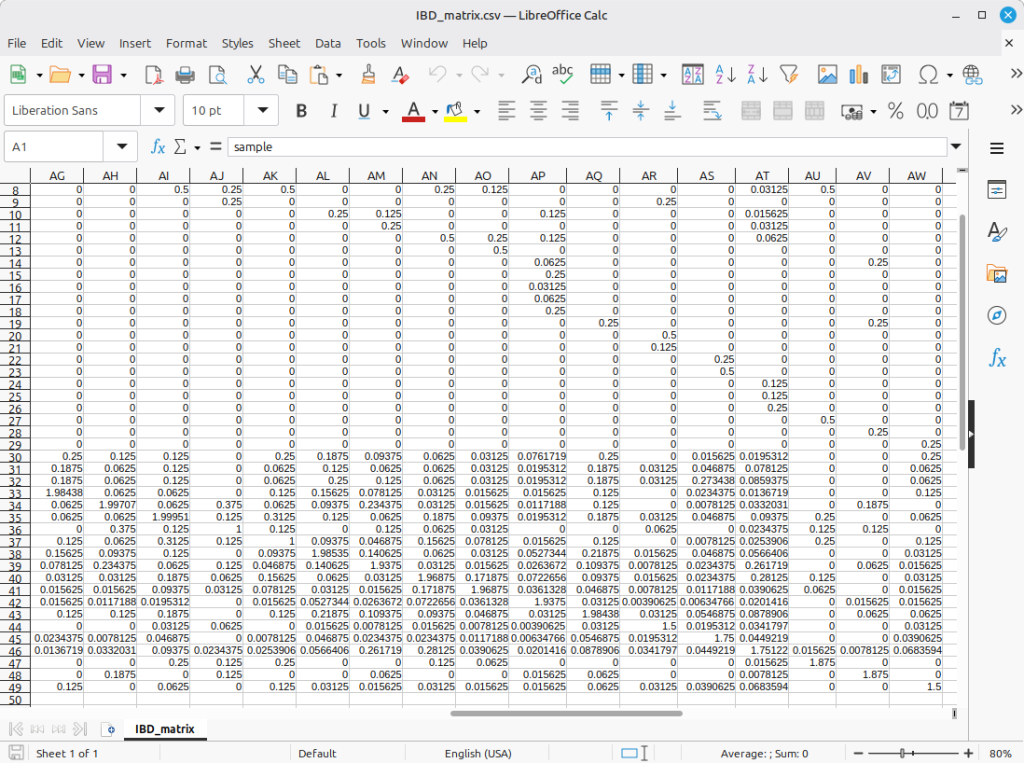
(Figure 2)
In “Pedigree editor”, you can freely edit and save pedigree records. Each time it is saved, a saved time point will be added to “Pedigree history”. This acts as a safeguard to prevent accidental data loss; if you mistakenly delete records and then save, you can simply restore your work by reverting to a specific time point before the error occurred. (See Figure 3)
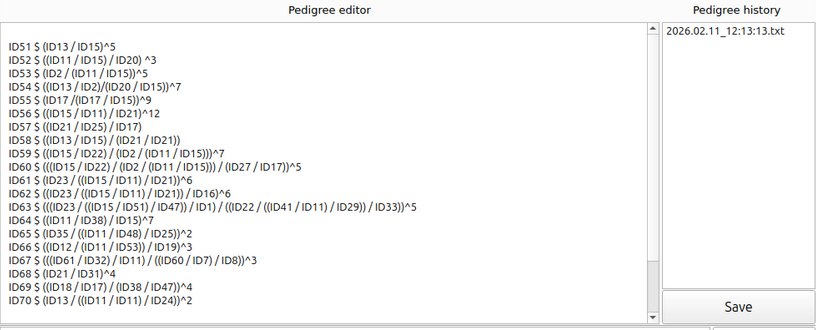
(Figure 3)
As aforementioned, the pedigree K matrix can be used to simulate a future kinship matrix by expanding a genomic K matrix based on scheduled pedigree (parental mating) records. If this is the case, make sure to check the check box labeled with “Use”, followed by uploading an existing genomic kinshp matrix as shwon in Figure 4.

(Figure 4)
Figure 5 shows an example of a simulated kinship matrix constructed from both a genomic kinship matrix and pedigree records.
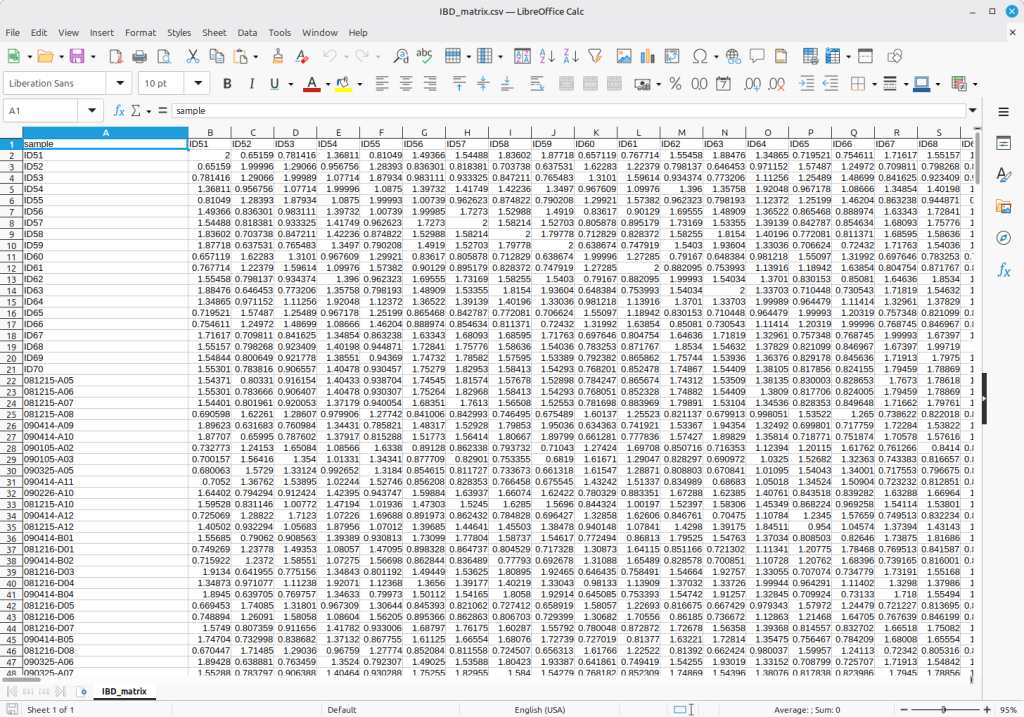
(Figure 5)